scDesignPop
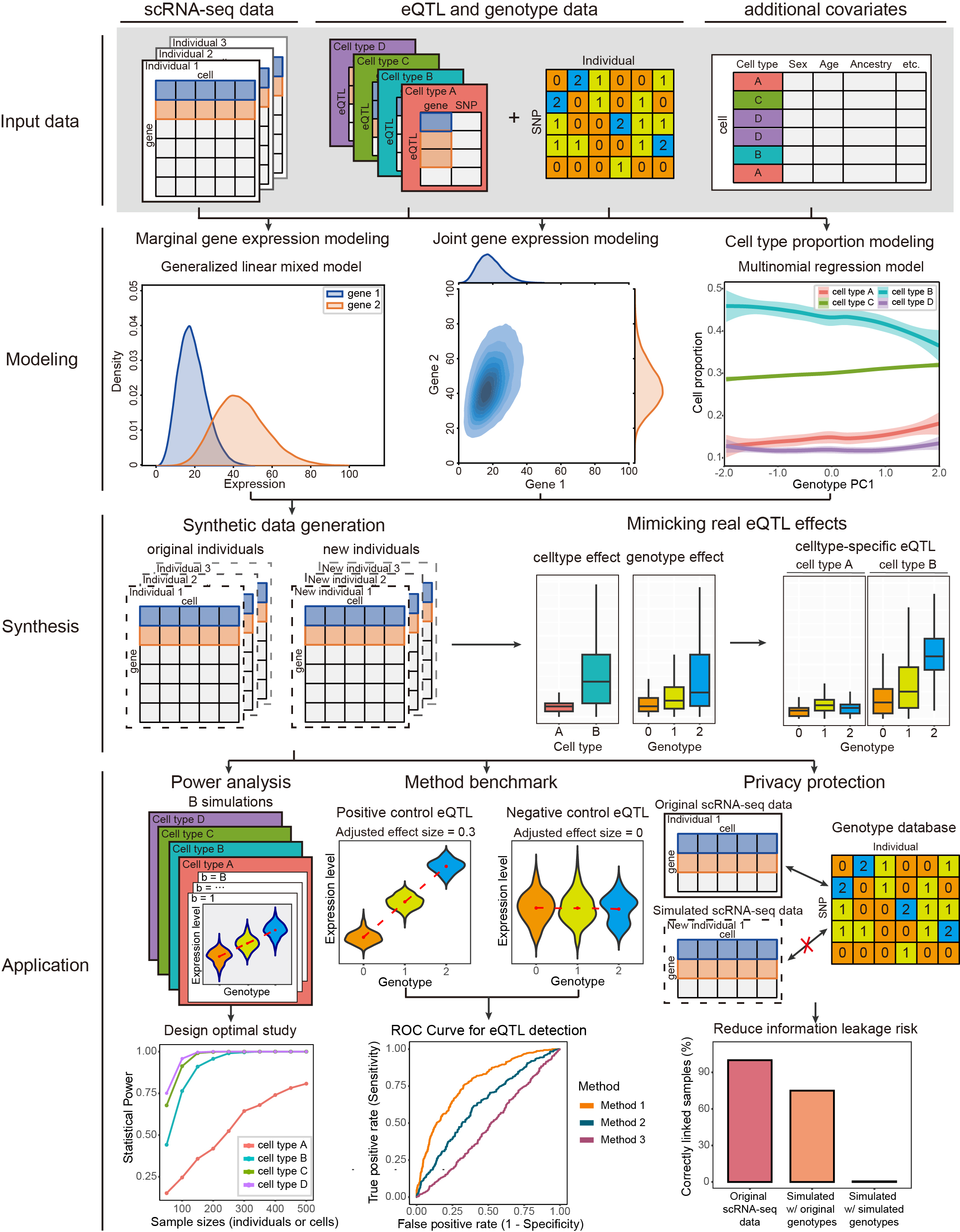
scDesignPop is a simulator for population-scale single-cell RNA-sequencing (scRNA-seq) data. By incorporating eQTL effects from genotype data with other covariates, scDesignPop has several key applications:
- performing eQTL power analysis across cell types,
- benchmarking single-cell eQTL mapping methods with user-specified eQTLs,
- protecting genomic privacy by mitigating eQTL-based re-identification of individuals via linking attacks
Detailed tutorials that illustrate various functionalities of scDesignPop are available at this website. The following illustration figure summarizes the workflow of scDesignPop:
Installation
To install the key dependence of scDesignPop, we recommend:
You can install the development version of scDesignPop from GitHub with:
if(!require("remotes")){
install.packages("remotes")
}
remotes::install_github("chrisycd/scDesignPop")Tutorials
For tutorials, please check the website. The tutorials will demonstrate the applications of scDesignPop as follows:
Contact
Any questions or advice on scDesignPop are welcomed! Please report it on issues, or contact Chris Dong (cycd@g.ucla.edu) or Yihui Cen (yihuicen@g.ucla.edu).
Related Manuscripts
The original scDesignPop paper (TBD. Coming soon…)
-
The simulator for general single-cell counts: scDesign3
-
The predecessors of scDesign3
- scDesign: Li, W. V., & Li, J. J. (2019). A statistical simulator scDesign for rational scRNA-seq experimental design. Bioinformatics, 35(14), i41-i50.
- scDesign2: Sun, T., Song, D., Li, W. V., & Li, J. J. (2021). scDesign2: a transparent simulator that generates high-fidelity single-cell gene expression count data with gene correlations captured. Genome biology, 22(1), 1-37.
-
The simulator for single-cell multi-omics reads developed by our lab member Guanao Yan